Monday, December 10, 2007


Pentose phosphate pathway
From Wikipedia, the free encyclopedia
Jump to: navigation, search
The pentose phosphate pathway (also called Phosphogluconate Pathway, or Hexose Monophosphate Shunt [HMP shunt]) is a cytosolic process that serves to generate NADPH and the synthesis of pentose (5-carbon) sugars. There are two distinct phases in the pathway. The first is the oxidative phase, in which NADPH is generated, and the second is the non-oxidative synthesis of 5 carbon sugars. This pathway is an alternative to glycolysis. While it does involve oxidation of glucose, its primary role is anabolic rather than catabolic.
Contents
[hide]
* 1 Uses
* 2 Phases
o 2.1 Oxidative phase
o 2.2 Non-oxidative phase
* 3 See also
* 4 References
* 5 External links
[ ] Uses
The pathway is one of the three main ways the body creates molecules with reducing power, accounting for approximately 60% of NADPH production in humans.
One of the uses of NADPH in the cell is to prevent oxidative stress. It reduces the coenzyme glutathione which converts reactive H2O2 into H2O. If absent, the H2O2 would be converted to hydroxyl free radicals which can attack the cell.
It is also used to generate hydrogen peroxide for phagocytes.[1]
[ ] Phases
[ ] Oxidative phase
In this phase, two molecules of NADP+ are reduced to NADPH, utilizing the energy from the conversion of glucose-6-phosphate into ribulose 5-phosphate.
Oxidative phase of pentose phosphate pathway
Oxidative phase of pentose phosphate pathway
The entire set of reactions can be summarized as follows:
Reactants Products Enzyme Description
Glucose 6-phosphate + NADP+ → 6-phosphoglucono-δ-lactone + NADPH glucose 6-phosphate dehydrogenase Dehydrogenation. The hemiacetal hydroxyl group located on carbon 1 of glucose 6-phosphate is converted into a carbonyl group, generating a lactone, and in the process NADPH is generated.
6-phosphoglucono-δ-lactone + H2O → 6-phosphogluconate + H+ 6-phosphoglucolactonase Hydrolysis
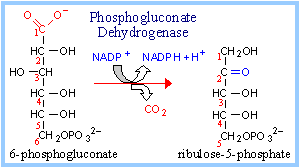
6-phosphogluconate + NADP+ → ribulose 5-phosphate + NADPH + CO2 6-phosphogluconate dehydrogenase Oxidative decarboxylation. NADP+ is the electron acceptor, generating another molecule of NADPH, a CO2, and ribulose 5-phosphate.
ribulose 5-phosphate ribose 5-phosphate Phosphopentose isomerase Isomerization. (Can also be considered part of nonoxidative phase)
The overall reaction for this process is:
Glucose 6-phosphate + 2 NADP+ + H2O → ribose 5-phosphate + 2 NADPH + 2 H+ + CO2
[ ] Non-oxidative phase
The pentose phosphate pathway's Nonoxidative phase
The pentose phosphate pathway's Nonoxidative phase
Reactants Products Enzymes
ribulose 5-phosphate → ribose 5-phosphate phosphopentose isomerase
ribulose 5-phosphate → xylulose 5-phosphate phosphopentose epimerase
xylulose 5-phosphate + ribose 5-phosphate → glyceraldehyde 3-phosphate + sedoheptulose 7-phosphate transketolase
sedoheptulose 7-phosphate + glyceraldehyde 3-phosphate → erythrose 4-phosphate + fructose 6-phosphate transaldolase
xylulose 5-phosphate + erythrose 4-phosphate → glyceraldehyde 3-phosphate + fructose 6-phosphate transketolase
GLUCONEOGENESIS







The vast majority of gluconeogenesis takes place in the liver and, to a smaller extent, in the cortex of kidneys. This process occurs during periods of fasting, starvation, or intense exercise and is highly endergonic. Gluconeogenesis is often associated with ketosis.
Contents
* 1 Entering the pathway
* 2 Pathway
* 3 Regulation
* 4 References
* 5 External links
Many 3- and 4-carbon substrates can enter the gluconeogenesis pathway. Lactate from anaerobic respiration in skeletal muscle is easily converted to pyruvate in the liver cells; this happens as part of the Cori cycle. However, the first designated substrate in the gluconeogenic pathway is pyruvate.
Oxaloacetate (an intermediate in the citric acid cycle) can also be used for gluconeogenesis. The gluconeogenic pathway can also generate glucose from amino acids, with the exception of lysine and leucine. Following removal of the amino group (by transamination or deamination) from the amino acid, the remaining carbon skeleton can enter gluconeogenesis directly (as pyruvate or oxaloacetate), or indirectly, e.g., via the citric acid cycle, converting α-ketoglutarate to oxaloacetate.
Fatty acids cannot be converted into glucose in animals, the exception being odd-chain fatty acids, which can yield propionyl CoA, a precursor for succinyl CoA. In plants, i.e., mainly in the seeds, the glyoxylate cycle allows conversion of fats into glucose, which is then used in the synthesis of complex carbohydrates, such as cellulose and glucans required for formation of new cell walls during germination. However, normally fatty acids are broken down into the two-carbon acetyl CoA, which is then used to fuel the citric acid cycle, and thus becomes unavailable to gluconeogensis. In contrast, glycerol, which is a part of all triacylglycerols, can be used in gluconeogenesis. In organisms in which glycerol is derived from glucose (e.g., humans and other mammals), glycerol is sometimes not considered a true gluconeogenic substrate, as it cannot be used to generate new glucose.
Pathway
* Gluconeogenesis is a pathway consisting of eleven enzyme-catalyzed reactions.
* Gluconeogenesis begins with the formation of oxaloacetate through carboxylation of pyruvate at the expense of one molecule of ATP, but is inhibited in the presence of high levels of ADP. This reaction is catalyzed by pyruvate carboxylase, which is stimulated by high levels of acetyl-CoA, i.e., when fatty acid oxidation is high in the liver.
* Oxaloacetate is then decarboxylated and simultaneously phosphorylated by phosphoenolpyruvate carboxykinase to produce phosphoenolpyruvate. One molecule of GTP is hydrolyzed to GDP in the course of this reaction. Oxaloacetate has to be reduced into malate using NADH in order to be transported out of the mitochondria. In the cytoplasm, malate is again oxidized to oxaloacetate using NAD+ where said reactions can occur.
* The next steps in the reaction are the same as reversed glycolysis. However fructose-1,6-bisphosphatase converts fructose-1,6-bisphosphate to fructose-6-phosphate. The purpose of this reaction is to overcome the large negative ΔG.
* Glucose-6-phosphate is formed from fructose-6-phosphate by phosphoglucoisomerase. Glucose-6-phosphate is used in other pathways. Free glucose is not generated automatically because glucose, unlike glucose-6-phosphate, tends to freely diffuse out of the cell. The reaction of actual glucose formation is carried out in the lumen of the endoplasmic reticulum. Here, glucose-6-phosphate is hydrolyzed by glucose-6-phosphatase, the last enzyme in gluconeogenesis, to produce glucose. Glucose is then shuttled into the cytosol by glucose transporters located in the membrane of the endoplasmic reticulum.
[ ] Regulation
Gluconeogenesis cannot be considered to be simply a reverse process of glycolysis, as the three irreversible steps in glycolysis are bypassed in gluconeogenesis. This is done to ensure that glycolysis and gluconeogenesis are not operating at the same time in the cell, making it a futile cycle. Therefore, glycolysis and gluconeogenesis follow reciprocal regulation, that is, cellular conditions, which inhibit glycolysis, may in turn activate gluconeogenesis.
Glucose-6-phosphate regulates the enzyme glucose-6-phosphatase in the lumen of ER by inducing its activity. In contrast, its accumulation will feed-back inhibit hexokinase in glycolysis. Once again, it follows the principle of reciprocal regulation.
The majority of the enzymes responsible for gluconeogenesis are found in the cytoplasm; the exceptions are mitochondrial pyruvate carboxylase, and, in animals, phosphoenolpyruvate carboxykinase. The latter exists as isozymes located in both the mitochondrion and the cytosol [1]. As there is no known mechanism to transport phosphoenolpyruvate from the mitochondrion into the cytosol, the cytosolic enzyme is believed to be the isozyme important for gluconeogeneis. The rate of gluconeogenesis is ultimately controlled by the action of a key enzyme, fructose-1,6-bisphosphatase, which is also regulated through signal tranduction by cAMP and its phosphorylation.
Most factors that regulate the activity of the gluconeogenesis pathway do so by inhibiting the activity or expression of key enzymes. However, both acetyl CoA and citrate activate gluconeogenesis enzymes (pyruvate carboxylase and fructose-1,6-bisphosphatase, respectively). Notably, acetyl-CoA and citrate also play inhibitory roles in pyruvate kinase activity in glycolysis.
Wednesday, December 5, 2007
krebs cycle

The citric acid cycle, also known as the tricarboxylic acid cycle (TCA cycle) or the Krebs cycle, (On rare occasions the citric acid cycle is known by a fourth name, the Szent-Györgyi-Krebs cycle) is a series of enzyme-catalysed chemical reactions of central importance in all living cells that use oxygen as part of cellular respiration. In eukaryotes, the citric acid cycle is located in the matrix of the mitochondrion. The components and reactions of the citric acid cycle were established by seminal work from both Albert Szent-Györgyi and Hans Krebs.
In aerobic organisms, the citric acid cycle is part of a metabolic pathway involved in the chemical conversion of carbohydrates, fats and proteins into carbon dioxide and water to generate a form of usable energy. Other relevant reactions in the pathway include those in glycolysis and pyruvate oxidation before the citric acid cycle, and oxidative phosphorylation after it. In addition, it provides precursors for many compounds including some amino acids and is therefore functional even in cells performing fermentation.
Contents
* 1 Overview
* 2 A simplified view of the process
* 3 Products
* 4 Regulation
* 5 Major metabolic pathways converging on the TCA cycle
* 6 See also
* 7 References
* 8 External links
Two carbons are oxidized to CO2, and the energy from these reactions is transferred to other metabolic processes by GTP (or ATP), and as electrons in NADH and FADH2. The NADH generated in the TCA cycle may later donate its electrons in oxidative phosphorylation to drive ATP synthesis; FADH2 is covalently attached to succinate dehydrogenase, an enzyme functioning both in the TCA cycle and the mitochondrial electron transfer chain in oxidative phosphorylation. FADH2 thereby facilitates transfer of electrons to coenzyme Q, an intermediate in the electron transfer chain.[1]
The citric acid cycle is continuously supplied new carbons in the form of acetyl-CoA, entering at step 1 below.[2]
S
t
e
p Substrates Products Enzyme Reaction type Comment
1 Oxaloacetate +
Acetyl CoA +
H2O Citrate +
CoA-SH Citrate synthase Aldol condensation rate limiting stage,
extends the 4C oxaloacetate to a 6C molecule
2 Citrate cis-Aconitate +
H2O Aconitase Dehydration reversible isomerisation
3 cis-Aconitate +
H2O Isocitrate Hydration
4 Isocitrate +
NAD+ Oxalosuccinate +
NADH + H + Isocitrate dehydrogenase Oxidation generates NADH (equivalent of 3 ATP)
5 Oxalosuccinate α-Ketoglutarate +
CO2 Decarboxylation irreversible stage,
generates a 5C molecule
6 α-Ketoglutarate +
NAD+ +
CoA-SH Succinyl-CoA +
NADH + H+ +
CO2 α-Ketoglutarate dehydrogenase Oxidative
decarboxylation generates NADH (equivalent of 3 ATP),
regenerates the 4C chain (CoA excluded)
7 Succinyl-CoA +
GDP + Pi Succinate +
CoA-SH +
GTP Succinyl-CoA synthetase substrate level phosphorylation or ADP->ATP,[1]
generates 1 ATP or equivalent
8 Succinate +
ubiquinone (Q) Fumarate +
ubiquinol (QH2) Succinate dehydrogenase Oxidation uses FAD as a prosthetic group (FAD->FADH2 in the first step of the reaction) in the enzyme,[1]
generates the equivalent of 2 ATP
9 Fumarate +
H2O L-Malate Fumarase H2O addition
(hydration)
10 L-Malate+
NAD+ Oxaloacetate +
NADH + H+ Malate dehydrogenase Oxidation generates NADH (equivalent of 3 ATP)
Mitochondria in animals including humans possess two succinyl-CoA synthetases, one that produces GTP from GDP, and another that produces ATP from ADP.[3] Plants have the type that produces ATP (ADP-forming succinyl-CoA synthetase).[2]
The GTP that is formed by GDP-forming succinyl-CoA synthetase may be utilized by nucleoside-diphosphate kinase to form ATP (the catalyzed reaction is GTP + ADP -> GDP + ATP).[1]
A simplified view of the process
* The citric acid cycle begins with acetyl-CoA transferring its two-carbon acetyl group to the four-carbon acceptor compound (oxaloacetate) to form a six-carbon compound (citrate).
* The citrate then goes through a series of chemical transformations, losing first one, then a second carboxyl group as CO2. The carbons lost as CO2 originate from what was oxaloacetate, not directly from acetyl-CoA. The carbons donated by acetyl-CoA become part of the oxaloacetate carbon backbone after the first turn of the citric acid cycle. Loss of the acetyl-CoA-donated carbons as CO2 requires several turns of the citric acid cycle. However, because of the role of the citric acid cycle in anabolism, they may not be lost since many TCA cycle intermediates are also used as precursors for the biosynthesis of other molecules.[4]
* Most of the energy made available by the oxidative steps of the cycle is transferred as energy-rich electrons to NAD+, forming NADH. For each acetyl group that enters the citric acid cycle, three molecules of NADH are produced.
* Electrons are also transferred to the electron acceptor FAD, forming FADH2.
* At the end of each cycle, the four-carbon oxaloacetate has been regenerated, and the cycle continues.
Products
Products of the first turn of the cycle are: one GTP, three NADH, one FADH2, two CO2.
Because two acetyl-CoA molecules are produced from each glucose molecule, two cycles are required per glucose molecule. Therefore, at the end of all cycles, the products are: two GTP, six NADH, two FADH2, and four CO2
Description Reactants Products
The sum of all reactions in the citric acid cycle is: Acetyl-CoA + 3 NAD+ + FAD + GDP + Pi + 2 H2O → CoA-SH + 3 NADH + 3 H+ + FADH2 + GTP + 2 CO2
Combining the reactions occurring during the pyruvate oxidation with those occurring during the citric acid cycle, the following overall pyruvate oxidation reaction is obtained: Pyruvic acid + 4 NAD+ + FAD + GDP + Pi + 2 H2O → 4 NADH + 4 H+ + FADH2 + GTP + 3 CO2
Combining the above reaction with the ones occurring in the course of glycolysis, the following overall glucose oxidation reaction (excluding reactions in the respiratory chain) is obtained: Glucose + 10 NAD+ + 2 FAD + 2 ADP + 2 GDP + 4 Pi + 2 H2O → 10 NADH + 10 H+ + 2 FADH2 + 2 ATP + 2 GTP + 6 CO2
(the above reactions are equilibrated if Pi represents the H2PO4- ion, ADP and GDP the ADP2- and GDP2- ions, respectively, and ATP and GTP the ATP3- and GTP3- ions, respectively).
Considering the future conversion of GTP to ATP and the maximum 32 ATP produced by the 10 NADH and the 2 FADH2 (see the theoretical yields for cellular respiration), it follows that each glucose molecule is able to produce a maximum of 32 ATP.
Regulation
Although pyruvate dehydrogenase is not technically a part of the citric acid cycle, its regulation is included here.
The regulation of the TCA cycle is largely determined by substrate availability and product inhibition. NADH, a product of all dehydrogenases in the TCA cycle with the exception of succinate dehydrogenase, inhibits pyruvate dehydrogenase, isocitrate dehydrogenase and α-ketoglutarate dehydrogenase, and also citrate synthase. Acetyl-CoA inhibits pyruvate dehydrogenase, while succinyl-CoA inhibits succinyl-CoA synthase and citrate synthase. When tested in vitro with TCA enzymes, ATP inhibits citrate synthase and α-ketoglutarate dehydrogenase; however, ATP levels do not change more than 10% in vivo between rest and vigorous exercise. There is no known allosteric mechanism that can account for large changes in reaction rate from an allosteric effector whose concentration changes less than 10% [5].
Calcium is used as a regulator. It activates pyruvate dehydrogenase, isocitrate dehydrogenase and α-ketoglutarate dehydrogenase.[6] This increases the reaction rate of many of the steps in the cycle, and therefore increases flux throughout the pathway.
Citrate is used for feedback inhibition, as it inhibits phosphofructokinase, an enzyme involved in glycolysis that catalyses formation of fructose 1,6-bisphosphate, a precursor of pyruvate. This prevents a constant high rate of flux when there is an accumulation of citrate and a decrease in substrate for the enzyme.
Recent work has demonstrated an important link between intermediates of the citric acid cycle and the regulation of hypoxia inducible factors (HIF). HIF plays a role in the regulation of oxygen haemostasis, and is a transcription factor which targets angiogenesis, vascular remodelling, glucose ulitisation, iron transport and apoptosis. HIF is synthesized consititutively and hydroxylation of at least one of two critical proline residues mediates their interation with the von Hippel Lindau E3 ubiquitin ligase complex which targets them for rapid degradation. This reaction is calalysed by prolyl 4-hydroxylases. Fumarate and succinate have been identified as potent inhibitors of prolyl hydroxylases thus leading to the stabilisation of HIF.[7]
Major metabolic pathways converging on the TCA cycle
Most of the body's catabolic pathways converge on the TCA cycle, as the diagram shows. Reactions that form intermediates of the TCA cycle in order to replenish them (especially during the scarcity of the intermediates) are called anaplerotic reactions.
The citric acid cycle is the third step in carbohydrate catabolism (the breakdown of sugars). Glycolysis breaks glucose (a six-carbon-molecule) down into pyruvate (a three-carbon molecule). In eukaryotes, pyruvate moves into the mitochondria. It is converted into acetyl-CoA by decarboxylation and enters the citric acid cycle.
In protein catabolism, proteins are broken down by protease enzymes into their constituent amino acids. The carbon backbone of these amino acids can become a source of energy by being converted to Acetyl-CoA and entering into the citric acid cycle.
In fat catabolism, triglycerides are hydrolyzed to break them into fatty acids and glycerol. In the liver the glycerol can be converted into glucose via dihydroxyacetone phosphate and glyceraldehyde-3-phosphate by way of gluconeogenesis. In many tissues, especially heart tissue, fatty acids are broken down through a process known as beta oxidation which results in acetyl-CoA which can be used in the citric acid cycle. Sometimes beta oxidation can yield propionyl CoA which can result in further glucose production by gluconeogenesis in the liver.
The citric acid cycle is always followed by oxidative phosphorylation. This process extracts the energy (as electrons) from NADH and FADH2, oxidizing them to NAD+ and FAD, respectively, so that the cycle can continue. Whereas the citric acid cycle does not use oxygen, oxidative phosphorylation does.
The total energy gained from the complete breakdown of one molecule of glucose by glycolysis, the citric acid cycle and oxidative phosphorylation equals about 36 ATP molecules. The citric acid cycle is called an amphibolic pathway because it participates in both catabolism and anabolism.
Sunday, December 2, 2007
nuclear receptor

Mechanism nuclear receptor action. This figure depicts the mechanism of a class I nuclear receptor (NR) which, in the absence of ligand, is located in the cytosol. Hormone binding triggers dissociation of heat shock proteins (HSP), dimerization, and translocation to the nucleus where it binds to a specific sequence of DNA known as a hormone response element (HRE). The nuclear receptor DNA complex in turn recruits other proteins that are responsible for transcription of downstream DNA into mRNA which is eventually translated into protein which results in a change in cell function.
Structures of selected endogenous nuclear receptor ligands and the name of the receptor that each binds to.
In the field of molecular biology, nuclear receptors are a class of proteins found within the interior of cells that are responsible for sensing the presence of hormones and certain other molecules. In response, these nuclear receptors affected by the hormones work in concert with other proteins to regulate the expression of specific genes, thereby regulating the mechanisms of the body (metabolism, developmental characteristics, homeostatic functions)
Nuclear receptors have the ability to directly bind to DNA and regulate the expression of adjacent genes, hence these receptors are classified as transcription factors.[1][2] The regulation of gene expression by nuclear receptors only happens when a ligand—a molecule which affects the receptor's behavior—is present. More specifically, ligand binding to a nuclear receptor results in a conformational change in the receptor which in turn activates the receptor resulting in up-regulation of gene expression.
A unique property of nuclear receptors which differentiate them from other classes of receptors is their ability to directly interact with and control the expression of genomic DNA. Consequently nuclear receptors play key roles in development and homeostasis of organisms. As discussed in more detail below, nuclear receptors may be classified either according to mechanism[3][4] or homology.[5][6]
Contents
[hide]
1 Ligands
2 Structure
3 Mechanism of action
3.1 Type I
3.2 Type II
3.3 Type III
3.4 Type IV
4 Coregulatory proteins
4.1 Coactivators
4.2 Corepressors
5 Agonism vs Antagonism
5.1 Agonists
5.2 Antagonists
5.3 Inverse agonists
5.4 Selective receptor modulators
6 Alternative mechanisms
6.1 Transrepression
6.2 Non-genomic
7 Family members
7.1 Subfamily 1: Thyroid Hormone Receptor-like
7.2 Subfamily 2: Retinoid X Receptor-like
7.3 Subfamily 3: Estrogen Receptor-like
7.4 Subfamily 4: Nerve Growth Factor IB-like
7.5 Subfamily 5: Steroidogenic Factor-like
7.6 Subfamily 6: Germ Cell Nuclear Factor-like
7.7 Subfamily 0: Miscellaneous
8 History of nuclear receptor
Ligands
Ligands that bind to and activate nuclear receptors include lipophilic substances such as endogenous hormones, vitamins A and D, and xenobiotic endocrine disruptors. Because the expression of a large number of genes is regulated by nuclear receptors, ligands that activate these receptors can have profound effects on the organism. Many of these regulated genes are associated with various diseases which explains why the molecular targets of approximately 13% of FDA approved drugs are nuclear receptors.[7]
A number of nuclear receptors, referred to as orphan receptors,[8] have no known (or at least generally agreed upon) endogenous ligands. Some of these receptors such as FXR, LXR, and PPAR bind a number of metabolic intermediates such as fatty acids, bile acids and/or sterols with relatively low affinity. These receptors hence may function as metabolic sensors. Other nuclear receptors, such as CAR and PXR appear to function as xenobiotic sensors up-regulating the expression of cytochrome P450 enzymes that metabolize these xenobiotics.[9]
[edit] Structure
Structural Organization of Nuclear Receptors
Top – Schematic 1D amino acid sequence of a nuclear receptor.
Bottom – 3D structures of the DBD (bound to DNA) and LBD (bound to hormone) regions of the nuclear receptor. The structures shown are of the estrogen receptor. Experimental structures of N-terminal domain (A/B), hinge region (D), and C-terminal domain (E) have not been determined therefore are represented by red, purple, and orange dashed lines respectively.
Nuclear receptors are modular in structure and contain the following domains:[10]
A-B) N-terminal regulatory domain: Contains the activation function 1 (AF-1) whose action is independent of the presence of ligand.[11] The transcriptional activation of AF-1 is normally very weak, but it does synergize with AF-2 (see below) to produce a more robust upregulation of gene expression. The A-B domain is highly variable in sequence between various nuclear receptors.
C) DNA-binding domain (DBD) (InterPro IPR001628): Highly conserved domain containing two zinc fingers (SCOP 57715) which binds to specific sequences of DNA called hormone response elements (HRE).
D) Hinge region: Thought to be a flexible domain which connects the DBD with the LBD. Influences intracellular trafficking and subcellular distribution.
E) Ligand binding domain (LBD) (InterPro IPR000536): Moderately conserved in sequence and highly conserved in structure between the various nuclear receptors. The structure of the LBD is referred to as an alpha helical sandwich fold (SCOP 48507) in which three anti parallel alpha helices (the "sandwich filling") are flanked by two alpha helices on one side and three on the other (the "bread"). The ligand binding cavity is within the interior of the LBD and just below three anti parallel alpha helical sandwich "filling". Along with the DBD, the LBD contributes to the dimerization interface of the receptor and in addition, binds coactivator and corepressor proteins. Contains the activation function 2 (AF-2) whose action is dependent on the presence of bound ligand.[11]
F) C-terminal domain: Variable in sequence between various nuclear receptors.
[edit] Mechanism of action
Mechanism nuclear receptor action. This figure depicts the mechanism of a class II nuclear receptor (NR) which, regardless of ligand binding status is located in the nucleus bound to DNA. For the purpose of illustration, the nuclear receptor shown here is thyroid hormone receptor (TR) heterodimerized to RXR. In the absence of ligand, TR is bound to corepressor protein. Ligand binding to TR causes a dissociation of corepressor and recruitment of coactivator protein which in turn recruit additional proteins such as RNA polymerase that are responsible for translation of downstream DNA into RNA and eventually protein which results in a change in cell function.
Nuclear receptors (NRs) may be classified into two broad classes according to their mechanism of action and subcellular distribution in the absence of ligand.
Small lipophilic substances such as natural hormones diffuse past the cell membrane and bind to nuclear receptors located in the cytosol (type I NR) or nucleus (type II NR) of the cell. This causes a change in the conformation of the receptor which depending on the mechanistic class (type I or II), triggers a number of down stream events that eventually results in up or down regulation of gene expression.
Accordingly, nuclear receptors may be subdivided into the following two mechanistic classes:[3][4]
[edit] Type I
Ligand binding to type I nuclear receptors in the cytosol (includes members of the NR subfamily 3) results in the dissociation of heat shock proteins, homo-dimerization, translocation (i.e., active transport) from the cytoplasm into the cell nucleus, and binding to specific sequences of DNA known as hormone response elements (HRE's). Type I nuclear receptors bind to HREs consisting of two half sites separated by a variable length of DNA and the second half site has a sequence inverted from the first (inverted repeat).
The nuclear receptor/DNA complex then recruits other proteins which transcribe DNA downstream from the HRE into messenger RNA and eventually protein which causes a change in cell function.
[edit] Type II
Type II receptors (principally NR subfamily 1) in contrast are retained in the nucleus regardless of the ligand binding status and in addition bind as hetero-dimers (usually with RXR) to DNA. In the absence of ligand, type II nuclear receptors are often complexed with corepressor proteins. Ligand binding to the nuclear receptor causes dissociation of corepressor and recruitment of coactivator proteins. Additional proteins including RNA polymerase are then recruited to the NR/DNA complex which translate DNA into messenger RNA.
[edit] Type III
Type III nuclear receptors (principally NR subfamily 2) are similar to type I receptors in that both classes bind to DNA has homodimers. However type III in contrast type I nuclear receptors bind to direct repeat instead of inverted repeat HREs.
[edit] Type IV
Type IV nuclear receptors bind either as monomers or dimers, but only a single DNA binding domain of the receptor binds to a single half site HRE. Examples of type IV receptors are found in most of the NR subfamilies.
[edit] Coregulatory proteins
Nuclear receptors bound to hormone response elements recruit a significant number of other proteins (referred to as transcription coregulators) which facilitate or inhibit the transcription of the associated target gene into mRNA.[12][13] The function of these coregulators are varied and include chromatin remodeling (making the target gene either more or less accessible to transcription) or a bridging function to stabilize the binding of other coregulatory proteins.
[edit] Coactivators
Binding of agonist ligands (see section below) to nuclear receptors induces a conformation of the receptor that preferentially binds coactivator proteins. These proteins often have an intrinsic histone acetyltransferase (HAT) activity which weakens the association of histones to DNA, and therefore promotes gene transcription.
[edit] Corepressors
Binding of antagonist ligands to nuclear receptors in contrast induces a conformation of the receptor that preferentially binds corepressor proteins. These proteins in turn recruit histone deacetylases (HDACs) which strengthens the association of histones to DNA, and therefore represses gene transcription.
[edit] Agonism vs Antagonism
Stuctural basis for the mechanism of nuclear receptor agonist and antagonist action.[14] The structures shown here are of the ligand binding domain (LBD) of the estrogen receptor (green cartoon diagram) complexed with either the agonist diethylstilbestrol (top, PDB 3ERD) or antagonist 4-hydroxytamoxifen (bottom, 3ERT). The ligands are depicted as space filling spheres (white = carbon, red = oxygen). When an agonist is bound to a nuclear receptor, the C-terminal alpha helix of the LDB (H12; light blue) is positioned such that a coactivator protein (red) can bind to the surface of the LBD. Shown here is just a small part of the coactivator protein, the so called NR box containing the LXXLL amino acid sequence motif.[15] Antagonists occupy the same ligand binding cavity of the nuclear receptor. However antagonist ligands in addition have a sidechain extension which sterically displaces H12 to occupy roughly the same postion in space as coactivators bind. Hence coactivator binding to the LBD is blocked.
Depending on the receptor involved, the chemical structure of the ligand and the tissue that is being affected, nuclear receptor ligands may display dramatically diverse effects ranging in a spectrum from agonism to antagonism to inverse agonism.[16]
[edit] Agonists
The activity of endogenous ligands (such as the hormones estradiol and testosterone) when bound to their cognate nuclear receptors is normally to upregulate gene expression. This stimulation of gene expression by the ligand is referred to as an agonist response. The agonistic effects of endogenous hormones can also be mimicked by certain synthetic ligands, for example, the glucocorticoid receptor antiiflammatory drug dexamethasone. Agonist ligands work by inducing a conformation of the receptor which favors coactivator binding (see upper half of the figure to the right).
[edit] Antagonists
Other synthetic nuclear receptor ligands have no apparent effect on gene transcription in the absence of endogenous ligand. However they block the effect of agonist through competitive binding to the same binding site in the nuclear receptor. These ligands are referred to as antagonists. An example of antagonistic nuclear receptor drug is mifepristone which binds to the glucocorticoid and progesterone receptors and therefore block the activity of the endogenous hormones cortisol and progesterone respectively. Antagonist ligands work by inducing a conformation of the receptor which prevents coactivator and promotes corepressor binding (see lower half of the figure to the right).
[edit] Inverse agonists
Finally, some nuclear receptors promote a low level of gene transcription in the absence of agonists (also referred to as basal or constitutive activity). Synthetic ligands which reduce this basal level of activity in nuclear receptors are known as inverse agonists.
[edit] Selective receptor modulators
A number of drugs that work through nuclear receptors display an agonist response in some tissue while an antagonistic response in other tissues. This behavior may have substantial benefits since it may allow retaining the desired beneficial therapeutic effects of a drug while minimizing undesirable side effects. Drugs with this mixed agonist/antagonist profile of action are referred to as selective receptor modulators (SRMs). Examples include Selective Estrogen Receptor Modulators (SERMs) and Selective Progesterone Receptor Modulators (SPRMs). The mechanism of action of SRMs may vary depending on the chemical structure of the ligand and the receptor involved, however it is thought that many SRMs work by promoting a conformation of the receptor that is closely balanced between agonism and antagonism. In tissues where the concentration of coactivator proteins is higher than corepressors, the equilibrium is shifted in the agonist direction. Conversely in tissues where corepressors dominate, the ligand behaves as an antagonist.[17]
[edit] Alternative mechanisms
[edit] Transrepression
The most common mechanism of nuclear receptor action involves direct binding of the nuclear receptor to a DNA hormone response element. This mechanism is referred to as transactivation. However some nuclear receptors not only have the ability to directly bind to DNA, but also to other transcription factors. This binding often results in deactivation of the second transcription factor in a process known as transrepresson.[18]
[edit] Non-genomic
The classical direct effects of nuclear receptors on gene regulation normally takes hours before a functional effect is seen in cells because of the large number of intermediate steps between nuclear receptor activation and changes in protein expression levels. However it has been observed that some effects from the application of hormones such as estrogen occur within minutes which is inconsistent with the classical mechanism nuclear receptor action. While the molecular target for these non-genomic effects of nuclear receptors has not been conclusively demonstrated, it has been hypothesized that there are variants of nuclear receptors which are membrane associated instead of being localized in the cytosol or nucleus. Furthermore these membrane associated receptors function through alternative signal transduction mechanisms not involving gene regulation.[19][20]
[edit] Family members
The following is a list of the 48 known human nuclear receptors[21] categorized according to sequence homology.[5][6] The list is organized as follows:
--------------------------------------------------------------------------------
Subfamily: name
Group: name (endogenous ligand if common to entire group)
Member: name (abbreviation; NRNC Symbol[5], gene) (endogenous ligand)
--------------------------------------------------------------------------------
Phylogenetic tree of human nuclear receptors
[edit] Subfamily 1: Thyroid Hormone Receptor-like
Group A: Thyroid hormone receptor (Thyroid hormone)
1: Thyroid hormone receptor-α (TRα; NR1A1, THRA)
2: Thyroid hormone receptor-β (TRβ; NR1A2, THRB)
Group B: Retinoic acid receptor (Vitamin A and related compounds)
1: Retinoic acid receptor-α (RARα; NR1B1, RARA)
2: Retinoic acid receptor-β (RARβ; NR1B2, RARB)
3: Retinoic acid receptor-γ (RARγ; NR1B3, RARG)
Group C: Peroxisome proliferator-activated receptor
1: Peroxisome proliferator-activated receptor-α (PPARα; NR1C1, PPARA)
2: Peroxisome proliferator-activated receptor-β/δ (PPARβ/δ; NR1C2, PPARD)
3: Peroxisome proliferator-activated receptor-γ (PPARγ; NR1C3, PPARG)
Group D: Rev-ErbA
1: Rev-ErbAα (Rev-ErbAα; NR1D1)
2: Rev-ErbAβ (Rev-ErbAβ; NR1D2)
Group F: RAR-related orphan receptor
1: RAR-related orphan receptor-α (RORα; NR1F1, RORA)
2: RAR-related orphan receptor-β (RORβ; NR1F2, RORB)
3: RAR-related orphan receptor-γ (RORγ; NR1F3, RORC)
Group H: Liver X receptor-like
3: Liver X receptor-α (LXRα; NR1H3)
2: Liver X receptor-β (LXRβ; NR1H2)
4: Farnesoid X receptor (FXR; NR1H4)
Group I: Vitamin D receptor-like
1: Vitamin D receptor (VDR; NR1I1, VDR) (vitamin D)
2: Pregnane X receptor (PXR; NR1I2)
3: Constitutive androstane receptor (CAR; NR1I3)
[edit] Subfamily 2: Retinoid X Receptor-like
Group A: Hepatocyte nuclear factor-4 (HNF4)
1: Hepatocyte nuclear factor-4-α (HNF4α; NR2A1, HNF4A)
2: Hepatocyte nuclear factor-4-γ (HNF4γ; NR2A2, HNF4G)
Group B: Retinoid X receptor (RXRα)
1: Retinoid X receptor-α (RXRα; NR2B1, RXRA)
2: Retinoid X receptor-β (RXRβ; NR2B2, RXRB)
3: Retinoid X receptor-γ (RXRγ; NR2B3, RXRG)
Group C: Testicular receptor
1: Testicular receptor 2 (TR2; NR2C1)
2: Testicular receptor 4 (TR4; NR2C2)
Group E: TLX/PNR
1: Human homologue of the Drosophila tailless gene (TLX; NR2E1)
3: Photoreceptor cell-specific nuclear receptor (PNR; NR2E3)
Group F: COUP/EAR
1: Chicken ovalbumin upstream promoter-transcription factor I (COUP-TFI; NR2F1)
2: Chicken ovalbumin upstream promoter-transcription factor II (COUP-TFII; NR2F2)
6: V-erbA-related gene|V-erbA-related (EAR-2; NR2F6)
[edit] Subfamily 3: Estrogen Receptor-like
See also steroid and sex hormone receptors
Group A: Estrogen receptor (Sex hormones: Estrogen)
1: Estrogen receptor-α (ERα; NR3A1, ESR1)
2: Estrogen receptor-β (ERβ; NR3A2, ESR2)
Group B: Estrogen related receptor
1: Estrogen-related receptor-α (ERRα; NR3B1, ESRRA)
2: Estrogen-related receptor-β (ERRβ; NR3B2, ESRRB)
3: Estrogen-related receptor-γ (ERRγ; NR3B3, ESRRG)
Group C: 3-Ketosteroid receptors
1: Glucocorticoid receptor (GR; NR3C1) (Cortisol)
2: Mineralocorticoid receptor (MR; NR3C2) (Aldosterone)
3: Progesterone receptor (PR; NR3C3, PGR) (Sex hormones: Progesterone)
4: Androgen receptor (AR; NR3C4, AR) (Sex hormones: Testosterone)
[edit] Subfamily 4: Nerve Growth Factor IB-like
Group A: NGFIB/NURR1/NOR1
1: Nerve Growth factor IB (NGFIB; NR4A1)
2: Nuclear receptor related 1 (NURR1; NR4A2)
3: Neuron-derived orphan receptor 1 (NOR1; NR4A3)
[edit] Subfamily 5: Steroidogenic Factor-like
Group A: SF1/LRH1
1: Steroidogenic factor 1 (SF1; NR5A1)
2: Liver receptor homolog-1 (LRH-1; NR5A2)
[edit] Subfamily 6: Germ Cell Nuclear Factor-like
Group A: GCNF
1: Germ cell nuclear factor (GCNF; NR6A1)
[edit] Subfamily 0: Miscellaneous
Group B: DAX/SHP
1: Dosage-sensitive sex reversal, adrenal hypoplasia critical region, on chromosome X, gene 1 (DAX1, NR0B1)
2: Small heterodimer partner (SHP; NR0B2)
Group C: Nuclear receptors with two DNA binding domains (2DBD-NR) (A novel subfamily)[22][23]
[edit] History of nuclear receptors
Below is a brief selection of key events in the history of nuclear receptor research.[24]
1905 - Ernest Starling coined the word hormone
1926 - Edward Calvin Kendall and Tadeus Reichstein isolated and determined the structures of cortisone and thyroxine
1929 - Adolf Butenandt and Edward Adelbert Doisy - independently isolated and determined the structure of estrogen
1961 - Elwood Jensen - isolated the estrogen receptor
1980s - cloning of the estrogen, glucocorticoid, and thyroid hormone receptors by Pierre Chambon, Ronald Evans, and Björn Vennström respectively
2004 - Pierre Chambon, Ronald Evans, and Elwood Jensen were awarded the Albert Lasker Award for Basic Medical Research, an award that frequently precedes a Nobel Prize in Medicine
Friday, November 23, 2007
glycolysis